Understanding marine biodiversity is paramount to environmental stewardship and the sustainability of ocean-related activities.
Environmental genomics provides comprehensive biological data and allows scientists to document all organisms without the need to directly observe or disturb them. Our DNA studies and monitoring programs produce higher resolution data and improved environmental insights while simplifying the current tedious methods for monitoring biodiversity. The technology allows scientists to collect and analyze a sample of eDNA from the environment, to identify the organisms present, and provide a detailed understanding of biodiversity. This is revolutionizing the biological and environmental sciences by bringing high-resolution data to the various levels of biological organization: from genomes to species to entire ecosystems. Our main pipelines are based on the eDNA metabarcoding approach:
The EnviroSeq® Genomics Workflow
Seven rigorous steps that yield repeatable, reproducible, and accurate results.

1. Program Design
A sophisticated process of planning, coordination and alignment of critical activities. Designing an eDNA survey is critical for successful implementation. It involves the planning, coordination and alignment of several activities, such as defining the scope, site selection, sampling strategy, marine logistics, choice of target DNA regions as biodiversity markers and selecting or designing DNA primer(s) for their amplification, selecting technology and depth of DNA sequencing, selecting bioinformatics workflow and other factors to ultimately deliver a comprehensive report that is targeted to the client’s requirements.

2. Calibration
Understanding local conditions. Every environment is different, and it is sometimes necessary to perform pilot studies to understand eDNA dynamics in previously unstudied regions. Factors such as: water temperature, hydrology, turbidity, richness of biodiversity, and composition of species will all affect recommendations on how to conduct a successful eDNA program.

3. Sample
Collect environmental samples of interest such as seawater, sediments, soil, or air. Using eDNA’s standardized collection protocols, the retrieval of eDNA will have little or no impact on the ecosystem or sampled populations, unlike traditional sampling techniques such as trawling, as no direct interaction with organisms is required to collect eDNA samples. We have developed specific protocols to collect and store samples in various environmental conditions including ambient temperature preservation. And we have hardened our sampling and transport logistics internationally, even from the most remote locations.
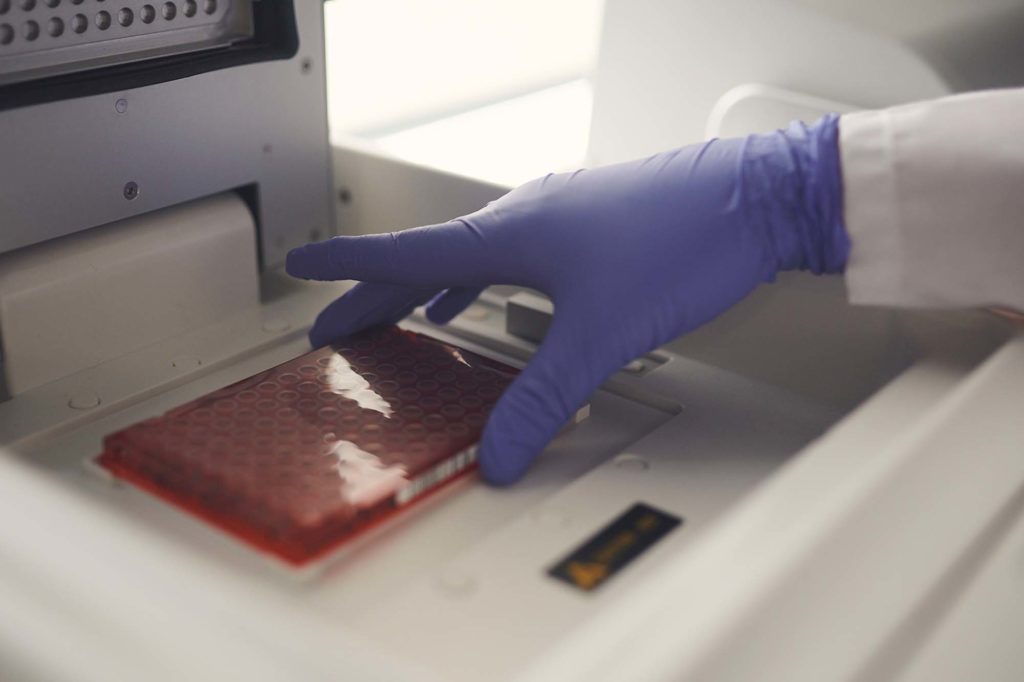

4. Extract environmental DNA to target species identification genes.
eDNA samples are extracted using eDNAtec’s standard operating protocols, and target gene regions are amplified using Polymerase Chain Reaction (PCR). This requires an understanding of which species (or groups of organisms) are of interest and may require the development of new “PCR primers” that will amplify DNA from the target groups.
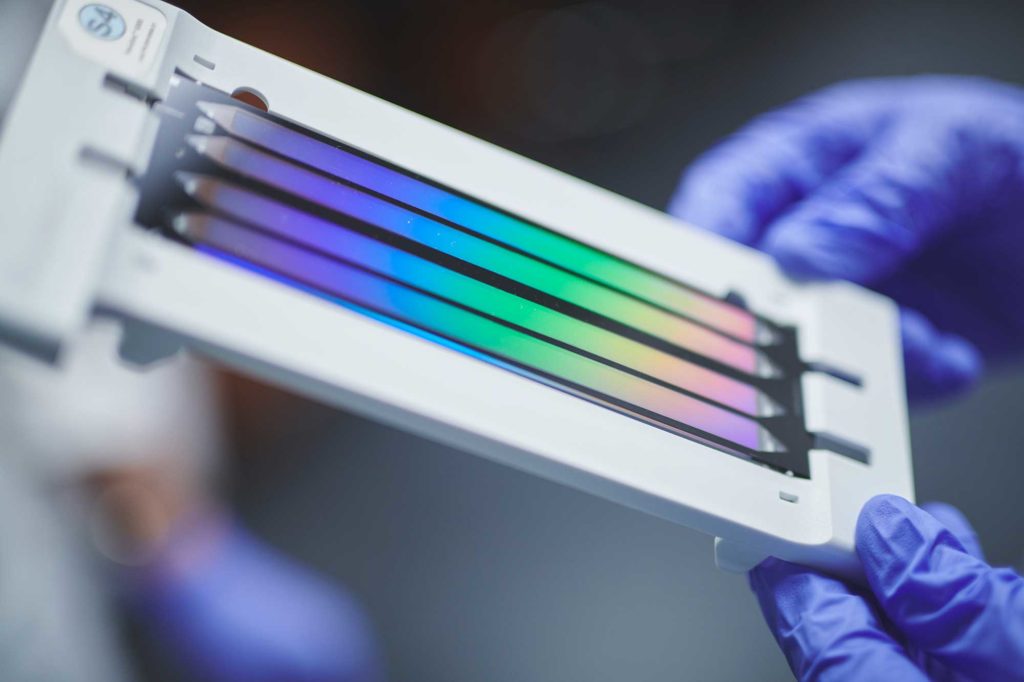

5. Read DNA sequences.
This is achieved using the most advanced high throughput technologies available, including the lllumina MiSeq and NovaSeq 6000 sequencing platforms. These are the same technologies used in advanced medical labs for various molecular diagnostic protocols.
6. Process sequence data using custom-designed bioinformatics software.
A typical survey will generate tens of millions or even billions of DNA sequences, which must pass through rigorous QC/QA checks and then be compared against a reference database of species to identify organisms.

7. Produce comprehensive biodiversity reports with actionable information.
eDNA analyses produce such a large amount of data with such high precision and accuracy that it enables views of the environment that were previously impossible. From high confidence predictive models about species distribution to reconstructing ecological food webs to the early detection of invasive species, eDNA technology empowers decision makers with better information.
